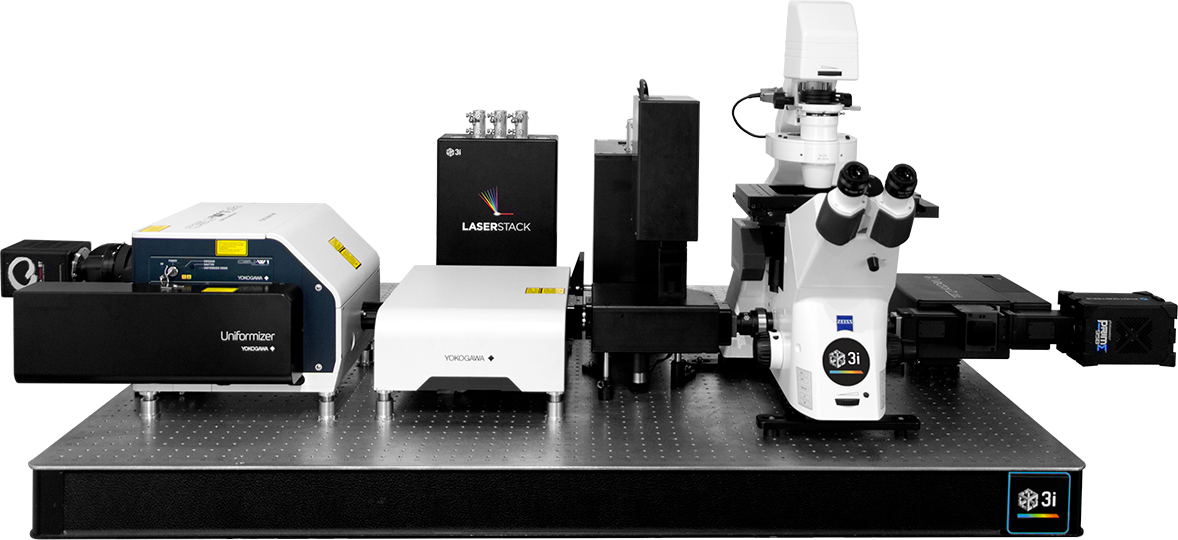



Up to four fiber outputs
Millisecond path switching
Environmental Control
Fully Automated Microscope




Computer generated holography
1-photon and 2-photon
Stage top and cage incubators
Temperature, gas and humidity
Motorized objective, condenser, path selection
Autofocus (Definite Focus 3)
PSF-optimized objectives
Laser ablation system
355nm and 532nm
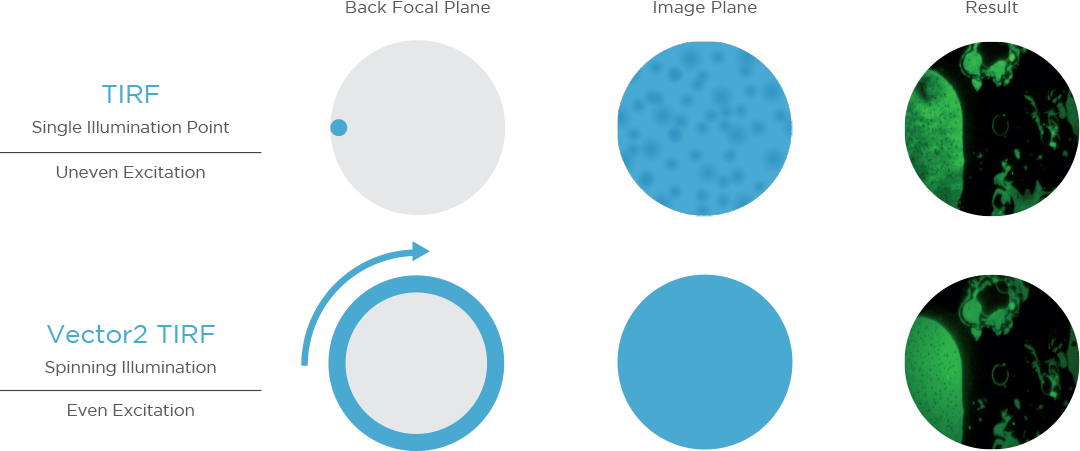
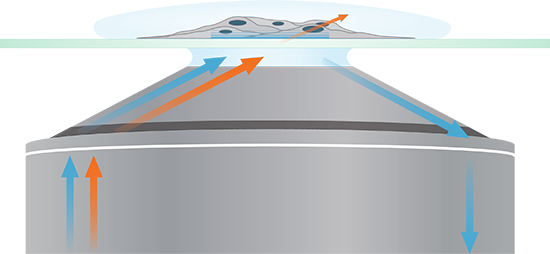
Spinning total internal reflection fluorescence
Modular high-speed X/Y scanner

Millisecond timing and trigger
Control of multiple devices
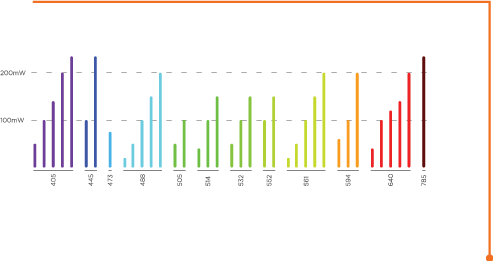
Modular laser combiner
Up to eight lasers

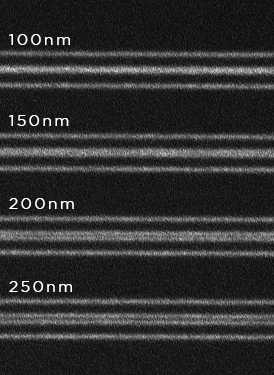
Live cell 3D confocal imaging
Super-resolution dual microlens disk

Path selection up to four ports
Sub-millisecond switch time

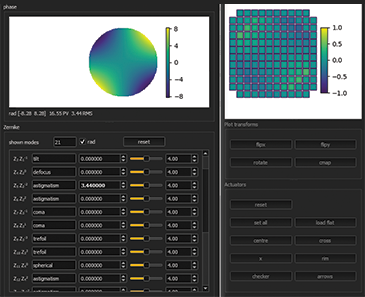
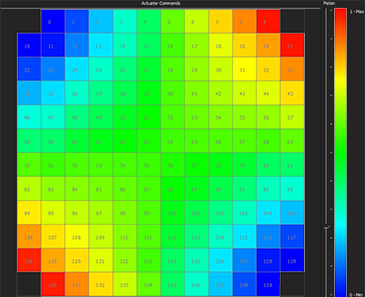
Dynamic correction of spherical aberration with depth





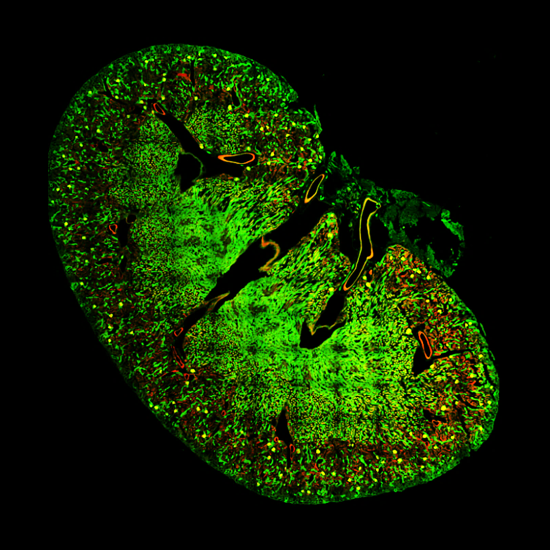
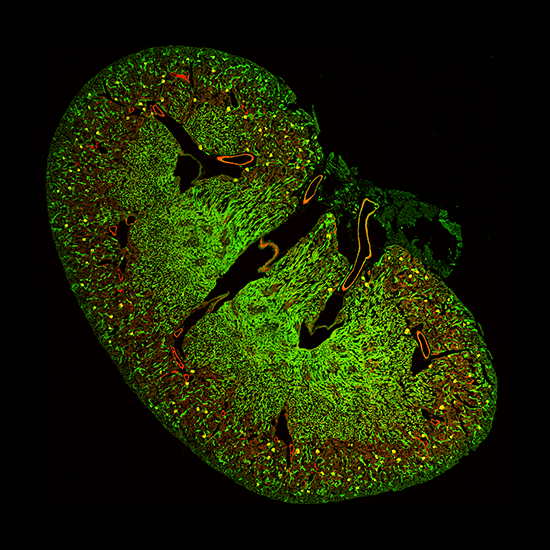
![100x Montage - Position 1 [25] Montage.Project Maximum Z](https://www.intelligent-imaging.com/wp-content/uploads/2022/03/100x-Montage-Position-1-25-Montage.Project-Maximum-Z.png)
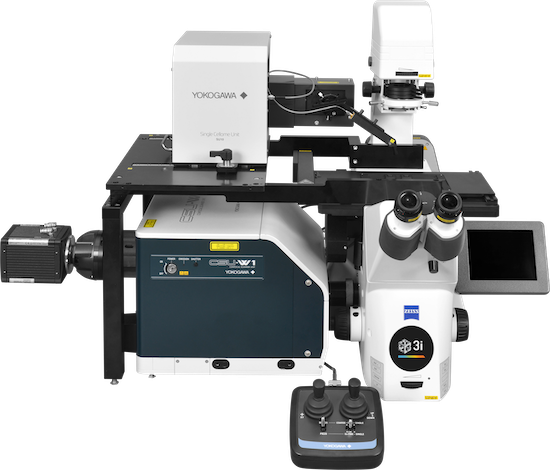
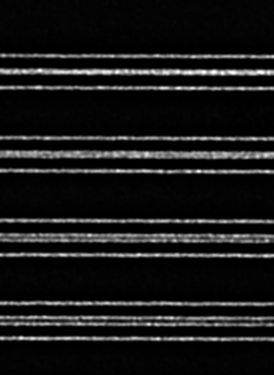
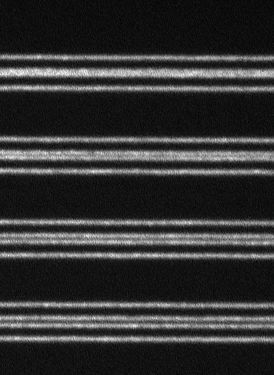





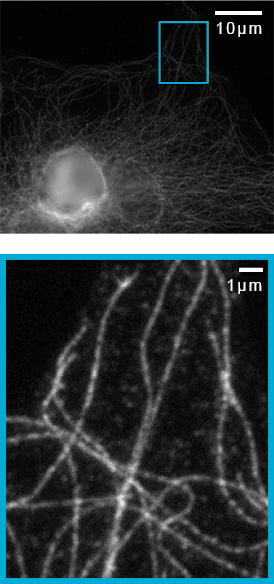
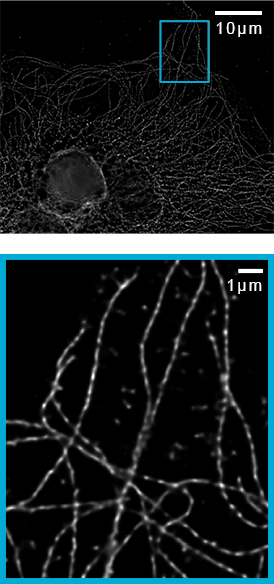
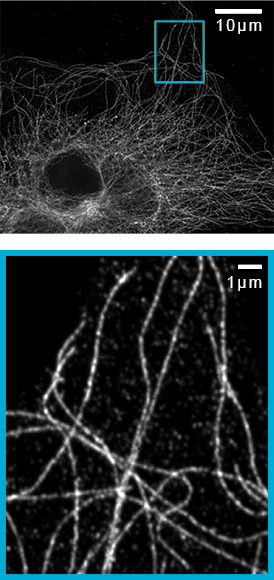
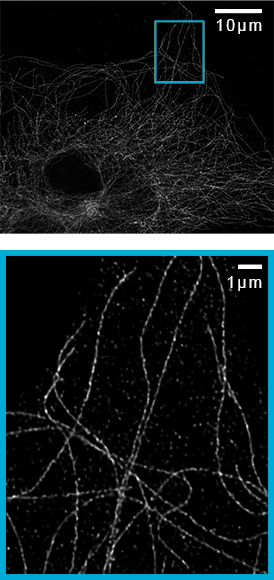
















![100x Montage 2.8x - Position 1 [49] Montage.Project Maximum Z](https://www.intelligent-imaging.com/wp-content/uploads/2022/04/100x-Montage-2.8x-Position-1-49-Montage.Project-Maximum-Z.png)